Rowan Zellers

I'm a researcher at OpenAI, working to build machines that understand humans and the world through language, vision, and sound. My pronouns are he/his.
I have a blog I sometimes update. You can also find me on Twitter at @rown, Mastodon at @rown@mastodon.social, or Bluesky at @rowan.bsky.social.
Publications
For all papers, visit my Semantic Scholar or Google Scholar pages, or .

ACL 2023
We introduce benchmarks that measure humor understanding, through explaining the jokes in New Yorker cartoons.
CVPR 2023
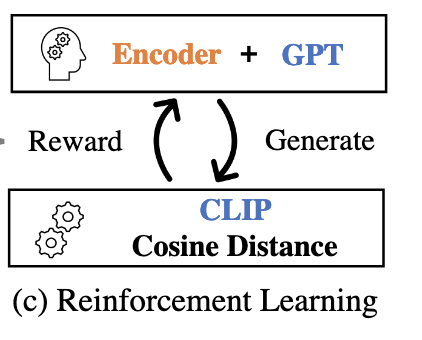
We use reinforcement learning to combine a vision model (CLIP) and a text-only language model, into a captioning model. The parameters of the language model are kept frozen, so the model can still perform zero-shot generalization, while also performing well on captioning.

CVPR 2022 Oral
website paper Press (VentureBeat) Press (GeekWire)
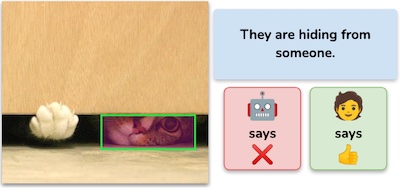
We introduce MERLOT Reserve, which learns about YouTube videos through all their modalities (audio, vision, and text). Learning from audio helps broadly -- even on single-image tasks like VCR. Our model learns state-of-the-art representations, that also transfer well to video-based tasks in a zero-shot setting.

arxiv 2022
We introduce a new dataset named Sherlock for studying Visual Abductive Reasoning: inferring likely context about the world beyond an image, given a localized clue. We operationalize this task through a ranking-based evaluation, where we find headroom for today's vision-and-language models.

NAACL 2022
The internet has lots of images paired with literal descriptions, but this pairing is less common for audio. We study using images as a "pivot" to learn strong audio-text representations, without paired data. In this zero-shot setting, our model obtains strong performance on a variety of sound description tasks.

NAACL 2022 Outstanding Paper (top 3)
We introduce a new constrained decoding strategy for text. The idea is to use lookahead heuristics to guide text towards satisfying provided constraints. It outperforms competitive baselines, particularly on tasks with complex constraints.

NeurIPS 2021 Oral (top 1% of submissions)
website paper Press (VentureBeat) Press (The Batch)
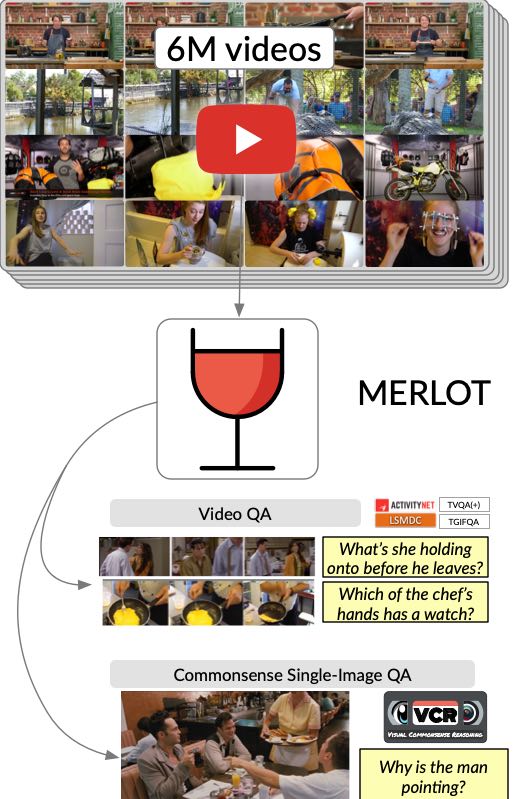
We introduce MERLOT, a model that learns multimodal script knowledge by watching millions of YouTube videos with transcribed speech -- in an entirely label-free, self-supervised manner. Our model not only learns to match images to temporally corresponding words, but also to contextualize what is happening globally over time. It shows a strong temporal understanding of visual events -- when finetuned, or when ordering visual stories zero-shot.
NeurIPS 2021 Outstanding paper (top 0.1%)
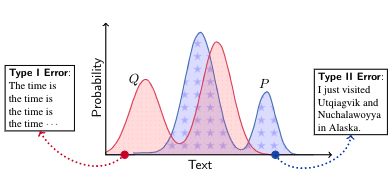
We introduce MAUVE, a new metric for evaluating open-ended text generation. MAUVE simultaneously measures 1) errors due to assigning high probability to unnatural language, and 2) errors due to assigning low probability to true language. Our evaluation corresponds with human judgments, and reveals that decoding algorithms like Nucleus Sampling score well in practice.

ACL 2021
We introduce a model named PIGLeT that learns physical commonsense understanding by interacting with the world, and uses this knowledge to ground language. This strategy works much better than modeling everything solely through linguistic form. When forecasting "what happens next" given an English sentence, our base-sized PIGLeT model outperforms text-to-text models that are 100x larger.
NAACL 2021
website paper Press (VentureBeat)
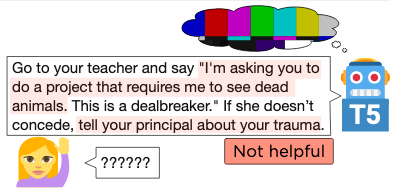
Big transformers do well on NLP benchmarks, but there's a gap between today's benchmarks and how humans use language. Our TuringAdvice benchmark narrows this gap, requiring models to write helpful language in response to a real-life situation. It reveals key flaws with today's models.

NAACL 2021
Constraints like "generate a recipe using the following ingredients" are tricky for language models to follow, even after finetuning. We introduce NeuroLogic Decoding, an algorithm that allows for the generation of high-quality text while satisfying constraints, allowing us to avoid the finetuning step completely.
ACL 2021
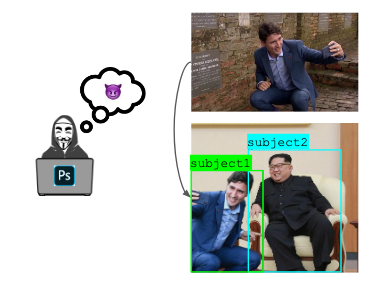
Many images online are edited in some way, but it is the intent that separates harmful edits like deepfakes from innocuous edits like an enhanced vacation photo. We introduce a new task and dataset for reasoning about why an image was edited, and a new model for the task.
NAACL 2021
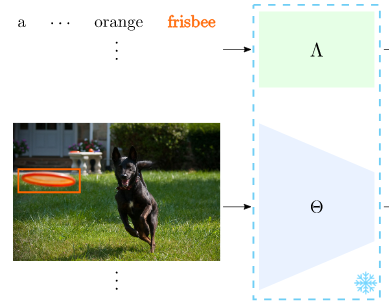
Today's best language understanding models are trained on massive amounts of purely textual data. We show that their representations have structural similarity to models trained on visual data, though this cross-modal linkage is far from perfect.

ICML 2020
Today's models achieve superhuman performance on benchmarks like ImageNet and Stanford Natural Language Inference (SNLI), but it is unclear whether they solved the underlying task, or rather overfitted to dataset biases. We study AFLite, an extension of Adversarial Filtering, to remove dataset biases. Filtering datasets like ImageNet makes them much harder for machines, while human performance remains high.

AAAI 2020 Oral (top 3%).
To apply eyeshadow without a brush, should I use a cotton swab or a toothpick? We introduce a benchmark for evaluating the ability for machines to reason about physical situations through natural language. Today's pretrained language models struggle, showing room for future reesearch.

NeurIPS 2019
website paper
Press: TechCrunch UW Allen School News New Scientist GeekWire AdWeek The New York Times The Washington Post AI2 Blog
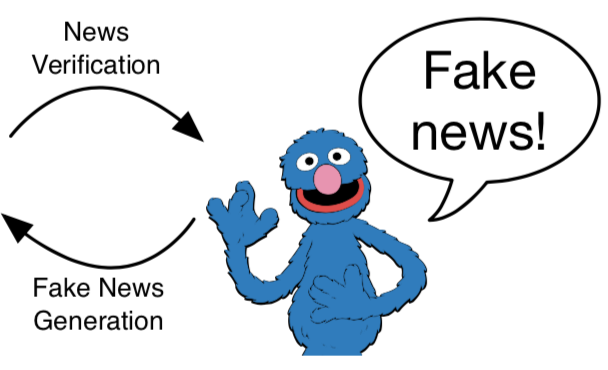
Can adversaries use state-of-the-art NLP models to generate "neural fake news"? We investigate the threat of machine-written propaganda that mimics the style of real news, through a model named Grover. Given a headline, Grover can generate the rest of the article. The best defense agianst Grover turns out to be Grover itself, demonstrating the importance of public release of strong generators.

ACL 2019
We show that commonsense inference still proves difficult for even state-of-the-art models, by presenting HellaSwag, a new challenge dataset. Though its questions are trivial for humans, even deep pretrained models (like BERT) struggle. The key insight is to scale up the length and complexity of the dataset examples towards a critical zone wherein generated text is ridiculous to humans, yet often misclassified by state-of-the-art models.

CVPR 2019 Oral (top 5%).
We formulate the new task of Visual Commonsense reasoning, where a model must not only answer challenging visual questions expressed in natural language: it must provide a rationale explaining why its answer is true. We introduce a new dataset, VCR, consisting of 290k multiple choice QA problems derived from 110k movie scenes.
EMNLP 2018
website paper Press (The New York Times)
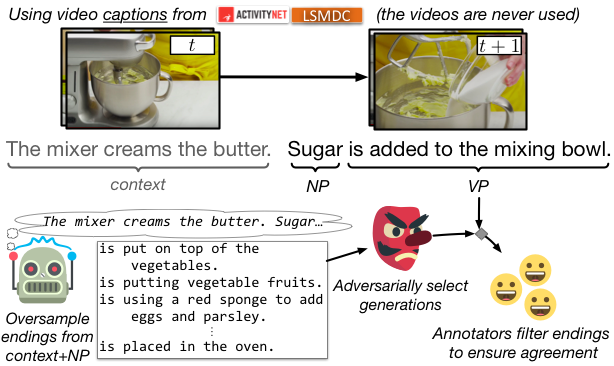
We release a new NLI dataset, SWAG, with 113k challenging multiple choice questions about grounded sitautions. To build this dataset, we present Adversarial Filtering (AF), which allows for data collection at scale while minimizing annotation artifacts.
CVPR 2018
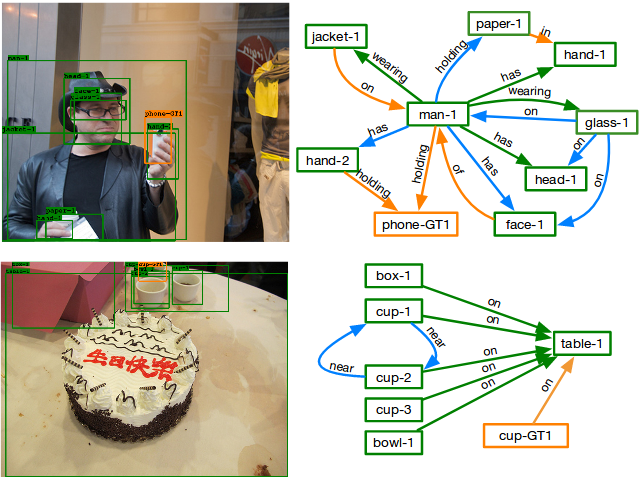
We study scene graph generation: building a graph where the nodes are objects and the edges are pairwise relationships. The visual world has many repeating structures (motifs). We built a model to capture them, which improves significantly over the prior staet-of-the-art.
EMNLP 2017
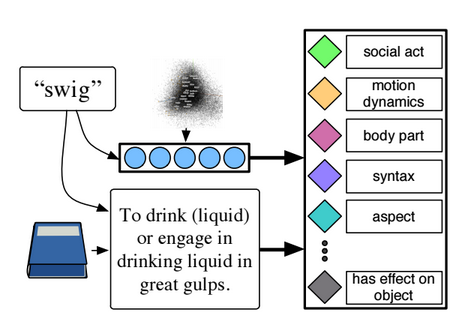
We investigate zero-shot multimodal learning where the topics of classification are verbs, not objects. We crowdsource verb attributes and build a model to learn them from unlabeled text, and dictionary definitions. We used these verb attributes to recognize actions in images.
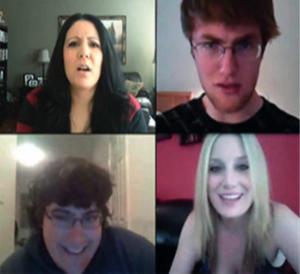
IEEE Intelligent Systems 2016
We provide a study of sentiment analysis applied on video data, not just text. We present a model that exploits the dynamics between gestures and verbal messages.
Recent Blog Posts
For more, visit my blog.
My Summer 2022 Europe adventure: Berlin, Amsterdam, Split, Mostar, Sarajevo, Belgrade, Budapest, Vienna.
March 03, 2024In Summer 2022, I went on an epic trip in Europe. I’m writing about it here, inspired by my friend Max’s travel blog.
Why I chose OpenAI over academia: reflections on the CS academic and industry job markets (part 2)
February 12, 2023At the end of my job search, I did something I totally wasn't expecting. I turned down all my academic job offers and signed the OpenAI offer instead.
Miscellaneous
- I graduated with my PhD from the University of Washington in Computer Science and Engineering and was part time at the Allen Institute for Artificial Intelligence. I worked with Yejin Choi and Ali Farhadi.
- During my PhD, I was supported by the NSF and ARCS Fellowships. Looking for tips on how to win the NSF fellowship? Check out my blog post here!
- Prior to UW, I graduated from Harvey Mudd College in 2016, where I majored in Computer Science and Math.
- As an undergrad, I worked with Louis-Philippe Morency on multimodal machine learning, and with Jacqueline Dresch on computational biology.
- Random things I like: rock climbing (especially bouldering!), running, biking, snowboarding, pottery, acro, art+design, urbanism, jazz guitar, Dominion, onebag travel...
- I have an Erdős number of 4 (Paul Erdős → Frank Harary → Chris Leary → Jacqueline Dresch → Me).
Contact
My email address is rowanz at cs.washington.edu. Note:
- If you have a question about getting code to work (for a paper I published), opening up an issue on my Github is usually better than email. Please try to provide sufficient detail so I can help you out!
- I often serve as a reviewer for open-access NLP, CV, and ML conferences and journals. I avoid reviewing for closed-access journals, or for conferences/journals that I haven't heard of, sorry!
